Difference between revisions of "GTRD"
Ivan Yevshin (Talk | contribs) (→Database statistics) |
Ivan Yevshin (Talk | contribs) (→Database statistics) |
||
| Line 32: | Line 32: | ||
! Object type !! Total count !! Per ChIP-seq experiment | ! Object type !! Total count !! Per ChIP-seq experiment | ||
|- | |- | ||
| − | | ChIP-seq reads || | + | | ChIP-seq reads || 183.8 × 10<sup>9</sup> || 36.2 × 10<sup>6</sup> |
|- | |- | ||
| − | | Reads aligned || | + | | Reads aligned || 146.9 × 10<sup>9</sup> || 28.9 × 10<sup>6</sup> |
|- | |- | ||
| − | | ChIP-seq peaks || | + | | ChIP-seq peaks || >100 × 10<sup>6</sup> || depends on peak caller |
|} | |} | ||
Revision as of 15:54, 26 August 2016
GTRD (Gene Transcription Regulation Database) is a database of transcription factor binding sites identified from ChIP-seq experiments that were systematically collected and uniformly processed using special workflow (pipeline) for BioUML platform.
Contents |
General info
GTRD (Gene Transcription Regulation Database) is a database of transcription factor binding sites identified from ChIP-seq experiments from:
Initial raw data were systematically collected and uniformly processed using specially developed workflow (pipeline) for BioUML platform:
- sequenced reads were aligned to reference genome using Bowtie;
- peaks were identified using MACS, SISSR, GEM and PICS peak callers
- peaks computed for the same TF and peak calling method, but different experiment conditions (e.g., cell line, treatment, etc.) were joined into clusters
- clusters for the same TF revealed by different peak calling methods were joined into meta-clusters
Learn more about GTRD build process
GTRD database is freely available for non-commercial organizations via web interface.
Database statistics
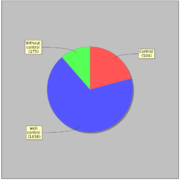
GTRD uses 5072 ChIP-seq experiments for 476 human and 257 mouse TFs that correspond to 542 TFClass classes. Most of ChIP-seq experiments (61%) have corresponding control experiment.General statistics:
| Object type | Total count | Per ChIP-seq experiment |
|---|---|---|
| ChIP-seq reads | 183.8 × 109 | 36.2 × 106 |
| Reads aligned | 146.9 × 109 | 28.9 × 106 |
| ChIP-seq peaks | >100 × 106 | depends on peak caller |
In average each TF has been measured in 9.37 ChIP-seq experiments, 54% of TFs have been measured in more than one experiment.
The ten most studied transcription factors are listed bellow:
| Transcription Factor | Number of ChIP-seq experiments |
|---|---|
| CTCF | 282 |
| AR | 117 |
| PU.1 | 103 |
| ERα | 92 |
| c-Myc | 79 |
| C/EBPβ | 74 |
| NF-κB p65 | 70 |
| GR | 53 |
| REST | 51 |
| GATA-1 | 51 |
Database structure
The metadata concerning GTRD is stored in MySQL tables.
Each ChIP-seq experiment has a row in 'chip_experiments' table, which assigns id and stores basic information about experiment. 'chip_experiments' table has following structure:
| Column | Description | Example value |
|---|---|---|
| id | Unique experiment identifier | EXP000489 |
| antibody | Antibody used in chromatin immunoprecipitation | sc-345 |
| tfClassId | Id in TFClass[1] database of target transcription factor, NULL for control experiments | 6.2.1.0.1 |
| cell_line | Studied cell line | HeLa S3 |
| specie | Species latin name | Homo sapiens |
| treatment | Cell treatment or conditions | IFN gamma |
| control_id | Id of control experiment, NULL for control experiments or experiments without control | EXP000490 |
The links to external databases stored in 'external_refs' table:
| Column | Description | Example values |
|---|---|---|
| id | Experiment identifier | EXP000489 |
| external_db | External database name | GEO or PUBMED or ENCODE or SRA |
| external_db_id | Identifier in external database | GSM320736 |
GTRD uses following object identifiers:
| Template | Object type | Example |
|---|---|---|
| EXPXXXXXX | ChIP-seq experiment | EXP000489 |
| READSXXXXXX | Collection of ChIP-seq reads | READS000770 |
| ALIGNSXXXXXX | Collection of read alignments | ALIGNS010001 |
| PEAKSXXXXXX | Collection of ChIP-seq peaks | PEAKS010000 |
The relationship between these objects is provided by 'hub' table:
| Column | Description | Example values |
|---|---|---|
| input | Input object identifier | READS000770 |
| input_type | Type of input object | ReadsGTRDType |
| output | Output object identifier | EXP000489 |
| output_type | Type of output object | ExperimentGTRDType |
ChIP-seq reads, alignments and peaks links to experiments with hub table in the following way:
| input | input_type | output | output_type |
|---|---|---|---|
| READS000770 | ReadsGTRDType | EXP000489 | ExperimentGTRDType |
| READS000771 | ReadsGTRDType | EXP000489 | ExperimentGTRDType |
| READS000772 | ReadsGTRDType | EXP000489 | ExperimentGTRDType |
| READS000773 | ReadsGTRDType | EXP000489 | ExperimentGTRDType |
| READS000774 | ReadsGTRDType | EXP000489 | ExperimentGTRDType |
| READS000775 | ReadsGTRDType | EXP000489 | ExperimentGTRDType |
| ALIGNS010001 | AlignmentsGTRDType | EXP000489 | ExperimentGTRDType |
| PEAKS010000 | PeaksGTRDType | EXP000489 | ExperimentGTRDType |
Web interface
| This page or section is a stub. Please add screenshots here! |
Web interface to GTRD is available here. It provides capabilities for searching and browsing GTRD.
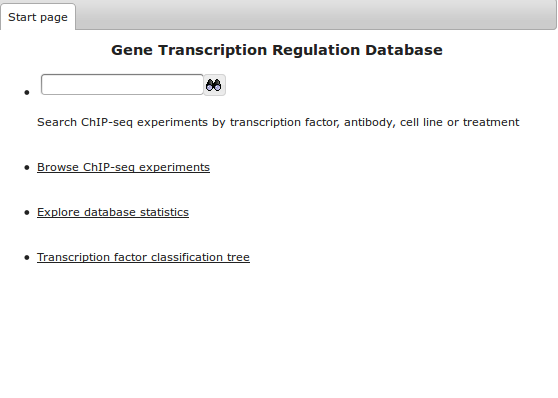
Start page
Start page contains search box and the links to browse all experiments in the repository tree, explore databases statistics and transcription factor classification tree.
Search capabilities
The ChIP-seq experiments contained in GTRD can be queried from the search box on the Start page. GTRD uses Lucene engine for indexing and quering ChIP-seq experiments that provides rich syntax for searching. The search can be performed by transcription factor(name or class), cell line, antibody or treatment/conditions.
For example, to search for STAT transcription factors enter stat* in the search field and press enter. You can restrict query to HeLa cells treated with interferon with following query:
tfTitle:stat* AND cellLine:hela AND treatment:IFN
The list of matching ChIP-seq experiments will appear in the 'Search result' tab. Select one of them to view detailed ChIP-seq experiment information in the information box. The information box also provides links to the experiment data: reads, alignments and peaks.
To view peaks or alignments click the link in the ChIP-seq experiment information box, the ![]() track will be opened as table. The
track will be opened as table. The ![]() track can be exported by pressing 'Export' button or opened in the genome browser by pressing 'Open as track' button in the general control panel.
track can be exported by pressing 'Export' button or opened in the genome browser by pressing 'Open as track' button in the general control panel.
Repository structure
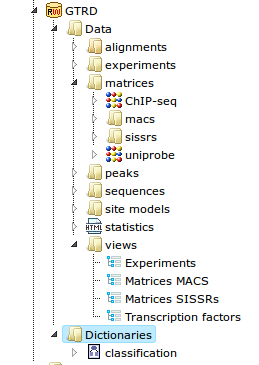
GTRD is organized in hierarchical Repository.
The GTRD/Data folder contains following items:
- experiments - ChIP-seq experiments metainformation
- sequences - Raw ChIP-seq reads in fastq.gz format
- alignments - ChIP-seq read alignments
- peaks - ChIP-seq peaks identified by MACS and SISSRs peak callers
- matrices - Position weight matrices for transcription factor binding sites
- site models - Models for recognition of transcription factor binding sites
- statistics - Summary GTRD statistics
- views - TFClass centric view of GTRD data avalable as
 tree-tables
tree-tables
GTRD/Dictionaries/classification is TFClass classification tree used by GTRD to reference transcription factors.